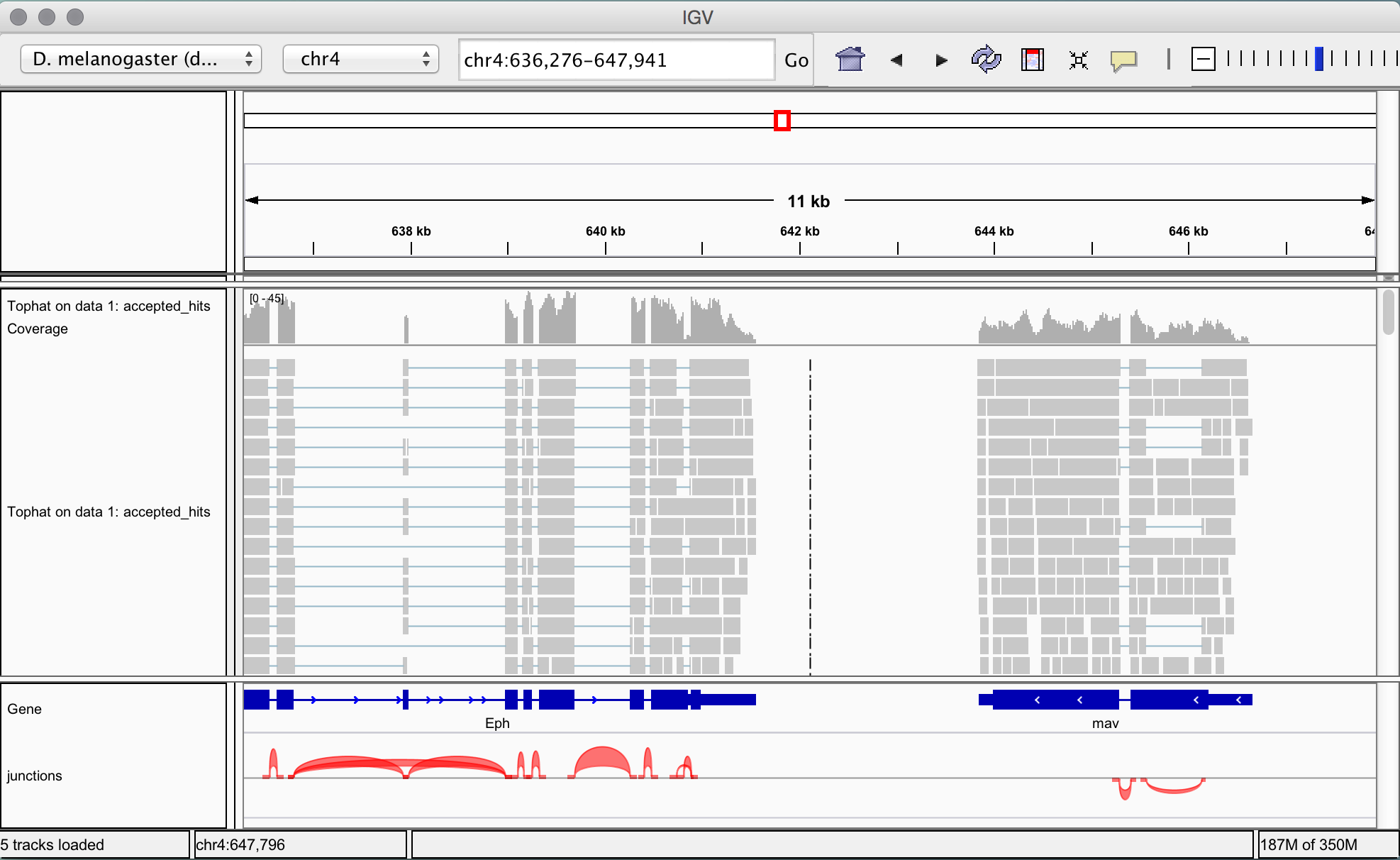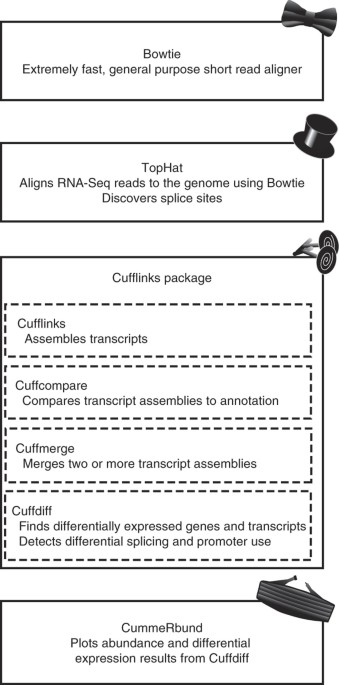
Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks | Nature Protocols
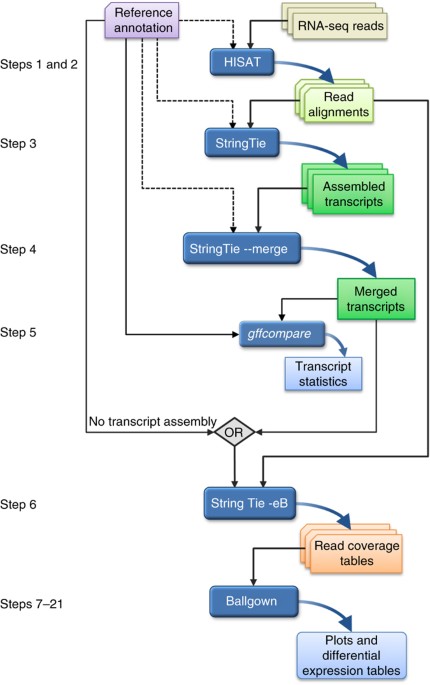
Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown | Nature Protocols
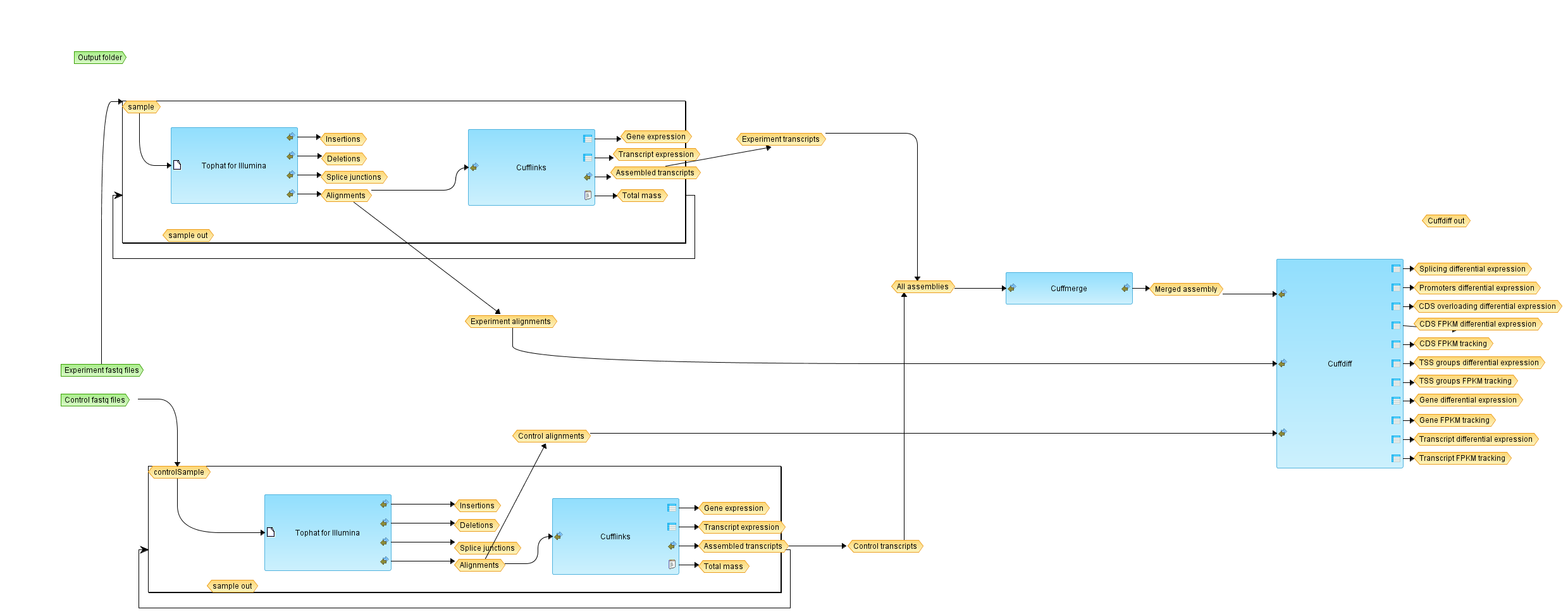
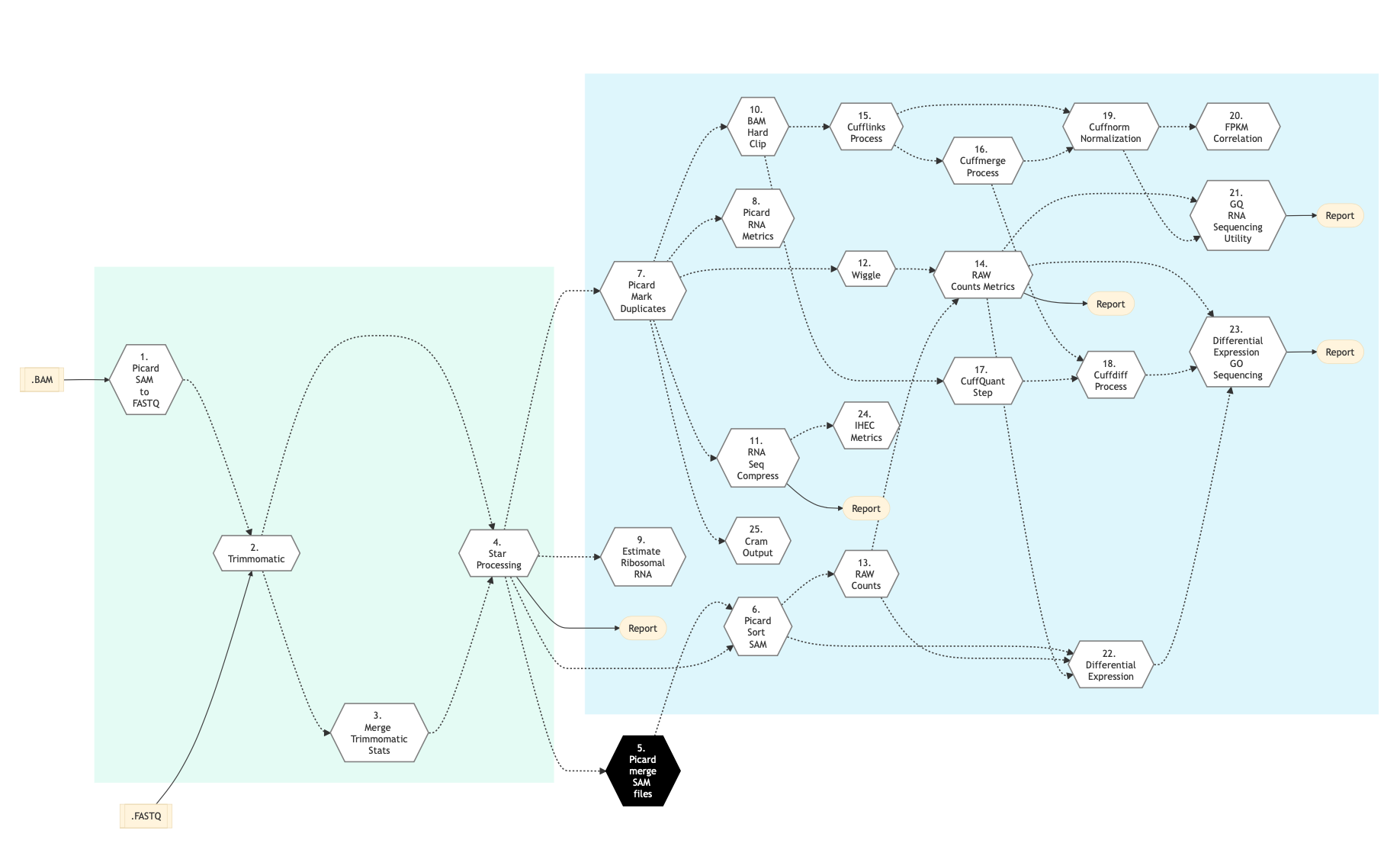
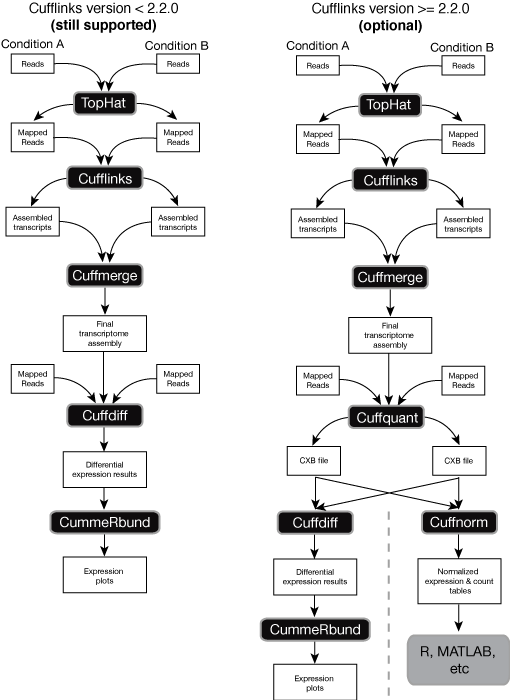
Quantification of RNA-seq with Cufflinks (with de-novo assembly) for FASTQ files (workflow) - BioUML platform

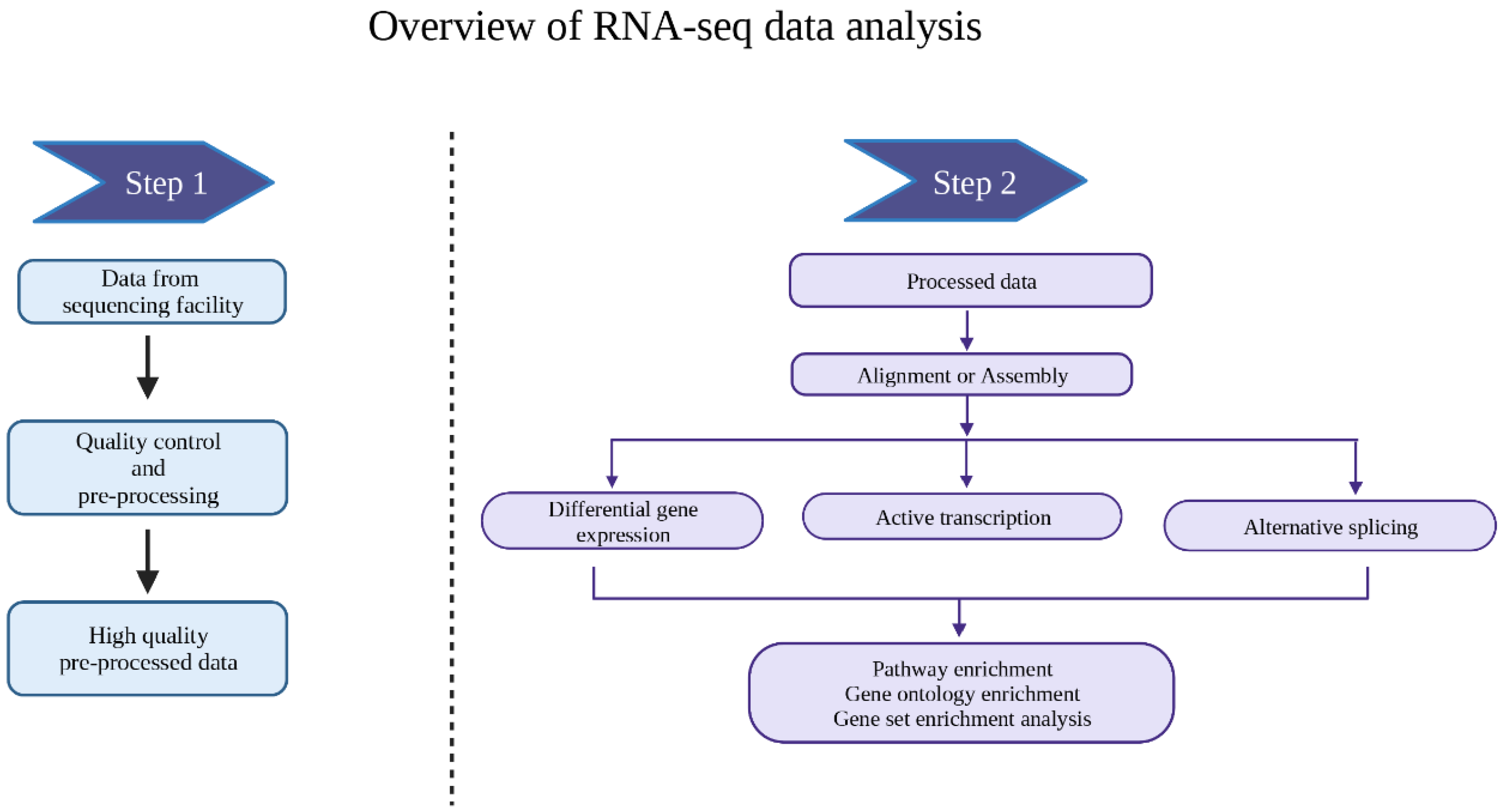
Principles of transcriptome analysis and gene expression quantification: an RNA‐seq tutorial - Wolf - 2013 - Molecular Ecology Resources - Wiley Online Library

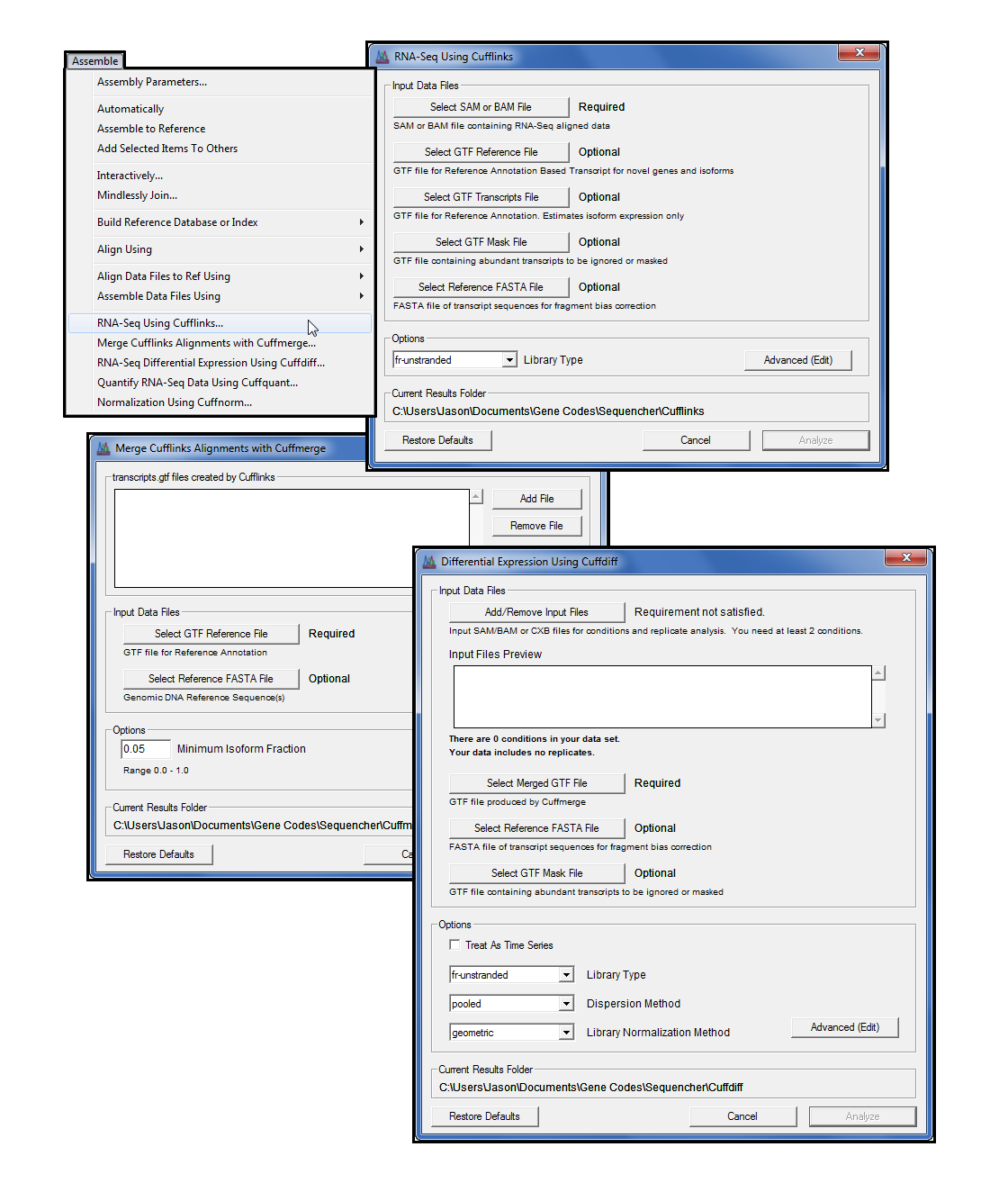
Differential Expression Analysis of RNA Sequencing Data using Cufflinks | by shilparaopradeep | Medium